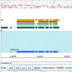
Bioinformatics defined
Prof. Allen Moore explains that bioinformatics can deal with a huge amount of genomic data, allowing researchers to explore complex relationships between many genes or genomes.
So bioinformatics has arisen because one of the problems is once we started sequencing genomes now we have an enormous amount of information. That information comes to us in the form of computer files with a lot of information that is not useful. So, bioinformatics is going through the information that you have and finding the useful bits. It’s very much mathematically based and using mathematical tools to start to push aside the parts that aren’t going to provide as much information and find those pieces of the data that are going to provide us with the most valuable information. So, bioinformatics arose because we have more data, more information, more pieces of DNA that are doing something than we can just look at. So in the old days, you might have one trait, find one gene, and put those two things together and know that that gene influences that trait. Now we are looking at very complicated traits that are influenced by lots an lots an lots of different parts of the genome and so we have to sift through all of the information we have to find out how that all fits together, and that’s basically what bioinformatics does. The other aspect of bioinformatics is that it uses what we already know to find related information inside something new, in a new project. So for example he might be interested in a gene that has been studied in mice and you want to know whether that gene or series of genes has similar influences in humans. You can compare mouse genome and human genome to see whether there are similar sorts of DNA exist in the human genome and then ask whether that then is associated with similar traits in humans.
bioinformatics, gene, genome, relationships, dna, technique, gene finding, mathematical tools, allen, moore
- ID: 2018
- Source: DNALC.G2C
- Download: MPEG 4 Video Theora Video Windows Media Video
Related Content
2020. Linkage versus association studies
Professor Allen Moore describes the differences between linkage and association studies, which are low- and high-resolution techniques used to search for candidate genes.
2014. What is DNA sequencing?
Professor Allen Moore explains that the DNA code is a long sequence made up of four bases (A,C,T, and G) and DNA sequencing is the processes of identifying the order in which they occur.
2016. Quantitative genetics studies
Professor Allen Moor explains that quantitative genetics is a technique for determining candidate genes for traits or disorders associated with multiple genes.
16086. Phylogenetic Tree; Macaulay
Phylogenetic tree modern humans mitochondrial genome mtDNA relationships eve bioinformatics
15290. Finding genes, Ewan Birney
Ewan Birney talks about finding genes.
2017. Linkage studies versus quantitative genetics
Professor Allen Moore outlines the differences between quantitative genetics and linkage studies. With quantitative genetics it is not necessary to begin with the physical DNA.
15291. Finding genes in the human genome, Ewan Birney
For the first draft of the genome sequence, both teams were working to identify the number of human genes. Here, Ewan Birney, a "numbers man" from the public genome project, explains how genes can be recognized and the data from the genome project used.
2015. Sequencing projects - new technology
Professor Allen Moore explains that since the beginning of the human genome project sequencing technology has become considerably cheaper and we now have sequences for many different organisms.
16990. DNA Subway (www.dnasubway.org)
The first educational product released by iPlant, DNA Subway (www.dnasubway.org) presents complex scientific tools and data in an intuitive and appealing interface, and makes high-level genome analysis broadly available to students and educators. "Riding"
2019. Expression analysis
Professor Allen Moore explains that expression analysis allows researchers to study what it is that the gene is making.