Animation 40: Living things share common genes.
Mike Wigler shows how all organisms share similar genes, called homologs.
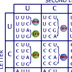
Hi, I’m Michael Wigler. Organisms share similar genes because they have inherited them from common ancestors. Even humans and yeast share similar genes! I was interested in genes involved in cancer, like the human ras gene. This gene contributes to uncontrolled cell growth and proliferation when mutated, for example, by the hydrocarbons in cigarette smoke. ras does not exist to cause cancer. We wanted to find out what it normally does so we looked for the same gene, or homolog, in yeast. If yeast have ras, we could use yeast instead of people to study the gene’s normal role. Nowadays, we can simply search for homologs on a large computer database, but in the early 1980s, we searched for homologs using DNA hybridization. Radioactive fragments of the human ras gene were used as probes to screen a gene library containing the entire yeast genome. We isolated clones that bound to the probe. Each of these clones contained a portion of the same gene — the yeast's ras gene. We sequenced the gene, deduced its amino acid sequence, and compared it to the human sequence. The yellow shading highlights amino acids that are identical in both proteins. As you can see, the two protein sequences are extremely similar. This means the gene has been conserved during the billion years since yeast and humans shared a common ancestor. All other eukaryotes also have this gene, and all the proteins resemble each other. The protein sequence has been conserved because it is essential for basic cell processes in these organisms. In fact, 33% of the yeast's genes are conserved in our own genome. When we compare the nucleotide sequences of the genes we also see similarity, but not as much. This is because the genetic code is redundant. Most amino acids are encoded by two to six different codons, so a change in one nucleotide does not necessarily change the amino acid. In the sequence below, 18 out of the 21 amino acid matches are encoded by different codons. Though we saw that the two genes are structural homologs — their amino acid sequences are very similar — that does not mean they do the same job. To see if the two genes perform the same function, we inserted the human gene into yeast cells. We started with a strain of yeast that was missing ras. The strain also lacked the leu gene, so we needed to supplement the medium with the amino acid leucine for the cells to grow. Then we added a plasmid to the yeast culture. The plasmid contained a functioning leu gene and a human ras gene that was under the control of a galactose promoter. We cut the plasmid and added it to the yeast culture. The plasmid integrated into the DNA of a few yeast cells. We isolated cells that had integrated the plasmid by spreading the yeast culture on a medium lacking leucine. Yeast that incorporated the plasmid and its leu gene survive and reproduce on this medium. Then we starved the transformed yeast so they would produce spores. Each yeast cell produces four spores encased in a capsule. We separated the spores onto different culture plates and watched them for germination. We knew from prior experience that yeast spores without ras would not germinate. None of the spores germinated on the glucose medium, because the galactose promoter needs galactose to turn on the human ras gene. On the galactose medium, the human ras gene was transcribed, and these spores germinated! Let's take a closer look to see why human ras can substitute for yeast ras and rescue the mutant yeast. (The amino acids are only labeled in the first row; the remaining residues are represented by dashes.) The red arrows point to a region of very high similarity in the first 80 residues. Here, the two proteins share 90% of their amino acids. Part of this region binds GTP during signal transduction, a job performed by all ras proteins – from yeast to fruit flies to humans. The GTP-binding domain has been conserved, because many substitutions here result in structural changes. Substituting the glycine at position 12, for example, pushes an adjacent branch away from the binding region. Click the hand to see the change. This mutation prevents the hydrolysis of GTP into GDP, and the protein activity can't turn off. In other words, the mutation turns a normal human ras gene into an oncogene — a gene that can cause uncontrolled cellular growth and cancer. 20% of human cancers carry this mutated ras. Other positions in this domain tolerate some change, as long as the substituted amino acid is similar. In yeast, position 11 is filled by glycine, the smallest amino acid, while in humans it is filled by alanine, the next smallest residue. There is no change in the protein's structure. We also noticed highly variable regions of the proteins. Some of these are not important for the protein's function. For example, the beginning of the ras proteins not only vary in the type of amino acid but also in the number of residues. Humans have three residues at the beginning of the protein, while yeast have ten! In human ras, amino acids in these positions can be replaced with unrelated residues without affecting the protein’s function. Hi, I’m Harold Varmus, and I’m Michael Bishop. Conserved genes tell the story of evolution as Darwin envisioned it: species inherit traits, with modification, from ancestral species. But genes are not always passed on in a linear fashion — organisms sometimes steal genes from one another. When we were studying cancer-causing retroviruses, we discovered that a virus had stolen a gene from a chicken! Remember, retroviruses carry their genetic information in RNA. A typical retrovirus has only three genes. A cancer-causing retrovirus has an additional gene. We started with two avian sarcoma viruses (ASV): the wild-type ASV that causes cancer in chickens and a mutant ASV that was missing the cancer-causing gene (src). Our first step was to isolate a probe for src. Using reverse transcriptase, we made single-stranded cDNAs from the wild-type virus and labeled them with radioactive hydrogen. We combined the cDNAs with single-stranded RNA we isolated from the mutant virus. Complementary sequences hybridized with each other to produce double-stranded molecules. Because the mutant virus lacked the src gene, the src cDNA was left in its single-stranded state. The single-stranded src cDNA we isolated became our probe. First, we combined the probe with single-stranded chicken DNA and let any complementary strands hybridize. We removed the single-stranded DNA with S1 nuclease, which only digest single-stranded DNA, and were left with radioactive hybrids. The probe had bound to the chicken's own src gene! But which came first, the chicken src or the viral src? We used the same probe to look for src genes in other animals. Birds, humans, mice and salmon had the gene, but invertebrates and bacteria did not. Further analysis showed that the viral src hybridized most completely with the chicken gene. It seemed unlikely that the virus gave src to all these animals when its src is most similar to the chicken's. We concluded the virus stole the gene from the chicken. Further proof came when we looked at the structure of the viral and chicken src genes. Hybridizing the viral src DNA to the chicken src revealed loops of introns in the chicken gene. The chicken src has introns, just like most other eukaryotic genes. The virus got rid of these introns when it captured the chicken gene in a process called transduction. The process depends on two viruses integrating their genomes into two places in the host. Sometimes, one of the viruses integrates next to an oncogene like src. If the downstream LTR – a sequence needed for viral integration – is deleted, the oncogene is copied into the viral progeny when the DNA is transcribed. The introns in the src gene are spliced out. Meanwhile, the RNA polymerase transcribes the second provirus. This normal virus RNA produces the viral capsid. Both RNAs get packaged into a capsid, where recombination generates a viral genome with both of the required LTRs; gag, env, and pol; and the src gene. This virus causes cancer when it infects a host, because the src gene, normally expressed at low levels, is controlled by a strong virus promoter. The src protein is overproduced, leading to uncontrolled growth.
nucleotide sequences, yeast genome, protein sequences, amino acid sequence, dna hybridization, common ancestor, wigler, cigarette smoke, protein sequence, amino acids, human sequence, uncontrolled cell growth, cell processes, codons, homolog, genetic code, hydrocarbons, shading, genes
- ID: 16834
- Source: DNALC.DNAFTB
Related Content
16854. Problem 40: Living things share common genes.
Find a cystic fibrosis protein function using bioinformatics to study homologs in model organisms.
555. Model Organisms
A human is a complicated organism, and it is considered unethical to do many kinds of experiments on human subjects. For these reasons, biologists often use simpler “model” organisms that are easy to keep and manipulate in the laboratory.
16514. Concept 23: A gene is a discrete sequence of DNA nucleotides.
Gene analysis take a giant leap using DNA sequencing.
16513. Problem 22: DNA words are three letters long.
Decode a protein.
16515. Animation 23: A gene is a discrete sequence of DNA nucleotides.
Fred Sanger outlines DNA sequencing.
1445. DNA
Because it contains the directions for assembling the components of the cell, DNA is often thought of as the "instruction book" for assembling life.
15501. Translation: RNA to protein, 3D animation with basic narration
3D animation of translation: RNA to protein.
16492. Problem 21: RNA is an intermediary between DNA and protein.
What happens in protein synthesis?
16494. Animation 22: DNA words are three letters long.
Several researchers crack the genetic code.
15481. Translation: RNA to protein, 3D animation with no audio
Translation: RNA to protein, 3D animation with no audio