DNA is packaged in a chromosome.
Roger Kornberg explains his work with Aaron Klug on histones, which bind DNA to form chromatin.
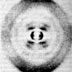
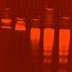
Hi, I'm Roger Kornberg. Aaron Klug and I were interested in a class of proteins called histones and how they interact with DNA. There are five different kinds of histones. Aaron Klug and I were interested in a class of proteins called histones and how they interact with DNA. There are five different kinds of histones. Histones bind to DNA to form the chromatin ("colored material") in the nucleus of higher cells. In non-dividing cells, the chromatin is dispersed throughout the nucleus. Histones bind to DNA to form the chromatin ("colored material") in the nucleus of higher cells. In non-dividing cells, the chromatin is dispersed throughout the nucleus. During prophase of cell division, the chromatin condenses into the visible structures we know as chromosomes. This electron micrograph shows a cell in metaphase; the chromosomes are lined up in the middle of the cell. The presence of histone proteins in the nucleus of higher cells was part of a debate in the 1940s about which molecule, DNA or proteins, is the hereditary material. Of course, DNA turned out to have that distinction. However, X-ray diffraction studies later showed that histones play an important role in providing structure for the DNA helix. Remember Maurice Wilkins? In 1964, he and Vittorio Luzzati noticed that chromatin has a repeating pattern with intervals of about 100 angstroms (1Å = 10 -10m). This repeat is different from the repeating patterns of DNA itself. Aaron Klug also saw similar X-ray diffraction patterns in chromatin. This repeat suggested that histones play a role in "packaging" DNA. I'm Dean Hewish. I'm Leigh Burgoyne. In 1973, while we were at Flinders University in South Australia, we got a result that supported the idea of DNA packaging. We isolated the enzyme DNA nuclease from rat liver cells and used it to digest chromatin. We electrophoresed the digested chromatin material ... ...and we saw a regular pattern of bands on the gel.This is a photo of the gel we ran in 1973. We figured out that the bands were multiples of the smallest size fragment, later determined to be about 200 base pairs (bp). So these repeated bands corresponded to 200 bp, 400 bp, 600 bp, 800 bp, and so on. We concluded that the histones are distributed evenly on the DNA and, at the points where they bind, protect the DNA from nuclease digestion. This is completely different from the digestion pattern of "naked" DNA without histones. Naked DNA digested with nuclease produces a smear of thousands of different-sized fragments. Based on the X-ray diffraction patterns and the nuclease experiments, chromatin was proposed to be DNA and the histone cores it wrapped around. The 200 bp repeat observed after nuclease digestion corresponds to 200 bp of DNA wrapped around each histone core. The 100Å measurement from X-ray diffraction patterns is the width of the histone core and the DNA. My colleagues and I did experiments that confirmed this model, and we also figured out the arrangement of the histones in the core. We individually purified the histones from the DNA. We found that H2A and H2B tend to stick together, as do H3 and H4. If we mixed the H2A/H2B complex with the H3/H4 complex, and then added naked DNA, we got the same X-ray pattern as for chromatin. More analysis revealed that each histone core has eight proteins -- two copies each of the H2A/H2B and H3/H4 complexes. This histone core with wrapped DNA is called a nucleosome. This is an electron micrograph of chromatin. The "string" is called the 10-nm fiber. The "beads" are the nucleosomes. But where is the H1 histone? It turns out H1 is not part of the histone core. Instead, it binds between nucleosomes to give even more structure to chromatin. H1 sits just outside of each nucleosome and interacts with the H1 in the next nucleosome. At higher salt concentrations, the 10-nm fiber is further compacted into the 30-nm fiber. The DNA helix is already twisted. By adding twists to make these nucleosomes and solenoid structures, the DNA is supercoiled. Even more organization is involved in maintaining the condensed chromosome. Loops of DNA are attached to a protein scaffold made up of several non-histone proteins. This scaffold maintains the shape of a chromosome -- even in the absence of histones. Chromosomes are really one continuous piece of DNA. In this electron micrograph you can see the DNA strand from one chromosome after the histones have been removed. Up to six feet of DNA is packaged to fit into the nucleus of one cell. The DNA is first wrapped around histone cores to form nucleosomes and the 10-nm fiber. The 10-nm fiber is further coiled into the 30-nm fiber, where six nucleosomes make one turn. The 30-nm fiber is then looped onto protein scaffolds when chromosomes condense.
x-ray diffraction, roger kornberg, electron micrograph, aaron klug, histone, maurice wilkins, vittorio luzzati, dean hewish, leigh burgoyne, dna, chromatin, prophase, metaphase, chromosome, electron micrograph, dna packaging, nuclease, nucleosome, gel
- ID: 16627
- Source: DNALC.DNAFTB
Related Content
15887. DNA packaging
DNA is coiled around proteins and packaged as chromatin within the nucleus of cells. Aaron Klug and Roger Kornberg figured out the structure of chromatin. It has been proposed that the coiling (or rather uncoiling) of DNA is a way of controlling the pro
16638. Gallery 29: Chromosome with histone stripped
Electron micrograph of the DNA and the protein scaffold left over from one chromosome (insert) with all the histone stripped out.
16636. Gallery 29: Electron micrograph of chromatin
Electron micrograph of the 10-nm fiber.
15875. Wilkins' X-ray
Maurice Wilkins obtained some of the first X-ray diffraction patterns of DNA from which dimensions could be calculated.
16637. Gallery 29: Electron micrograph of chromatin (1)
Electron micrograph of the 30-nm fiber.
15337. Use of X-ray crstallography to prove that DNA is crystalline, Maurice Wilkins
Maurice Wilkins talks about obtaining an X-ray diffraction pattern.
16644. Biography 29: Roger Kornberg (1947 - )
In 1974, Roger Kornberg worked out the importance of histones to chromatin structure.
16635. Gallery 29: Chromatin digested by nuclease
Photo of chromatin digested by nuclease, from Hewish and Burgoyne's 1973 experiment.
16034. Roger Kornberg, 1974
A chromosome is a package for DNA.
15484. DNA packaging, 3D animation with sound effects only
DNA packaging, 3D animation with sound effects only