Animation 38: Development balances cell growth and death.
Leland Hartwell describes how cells regulate the timing of growth and cell division. Bob Horvitz and Mike Hengartner explain control mechanisms for cell death.
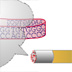
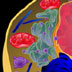
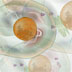
Hi, I'm Leland Hartwell. I was one of the first to use yeast cells as a model system to study biological problems. I was interested in how cells regulate the timing of growth and cell division. Yeasts are single-celled organisms that divide by budding. The process is the same as mitosis except that the nuclear membrane stays intact. The yeast cells in this photo express a fluorescent protein in their membranes, and you can see budding as well as non-budding yeast cells. Through my work with yeast cells, I found that all eukaryotic cells cycle through four different stages. A cell first has to grow and replenish its resources. This stage is called G1; the G stands for "gap." Next, the cell synthesizes DNA in preparation for cell division. This is the S stage. After DNA replication, the cell enters the second gap stage, G2, where it makes other proteins and cellular components necessary for cell division. M is the mitotic stage; the cell divides and the whole cycle repeats. Mitosis is dependent on the completion of all the other events in the other three stages. We isolated a number of yeast mutants that weren't able to complete this cycle. We called them cell division cycle (cdc) mutants. Some cdc mutants turned out to have defects in the replication machinery. For example, cdc9 has a defective DNA ligase. DNA ligase is a protein that knits DNA pieces together. Because DNA polymerase adds nucleotides in the 5' to 3' direction, only one strand, the leading strand, is replicated as a continuous piece of DNA. The other strand, the lagging strand, is actually made in short 5' to 3' streches called Okazaki fragments. DNA ligase then knits the pieces together to make a continuous strand. cdc9 mutants have no functioning DNA ligase. The Okazaki fragments are never knitted together. Of course, it made sense that some cdc mutants had defects in the replication machinery. But we also found other types of cdc mutants. Normally, when exposed to radiation, cells stop in G1 to repair DNA damage before continuing onto the S stage. We isolated a mutant, rad9, that did not stop in G1 after radiation treatment. It continued on and finished the cell cycle by dividing. We believed that rad9 cells could no longer detect DNA damage caused by radiation. Thus, rad9 mutants divide and finish the cell cycle even when their DNA is not "ready." We tested this idea by making a double mutant that was both rad9 and cdc9. Remember, we knew that cdc9 had a defect in DNA ligase, so the DNA of one of the newly synthesized strands was in pieces. The double mutant finished the cell cycle by dividing even though the lagging strand was in pieces. Thus, rad9 works by stopping cells from dividing when DNA replication is incomplete or when DNA damage has occurred. Thus, rad9 works by stopping cells from dividing when DNA replication is incomplete or when DNA damage has occurred. We called these genes checkpoint genes. They allow cells to proceed to the next stage of the cell cycle only when specific requirements, like DNA synthesis and replication, have been met. We also isolated genes that acted as checkpoints for the other cell stages. Many of these checkpoint genes are also found in other species. Hi, I'm Bob Horvitz. I'm Mike Hengartner. Lee Hartwell told you about control mechanisms for cell growth and division. We’re going to tell you about control mechanisms for cell death. Mike and I work on an organism called Caenorhabitis elegans (C. elegans). It's a non-parasitic, microscopic roundworm with only 959 cells. The fate and lineage of every cell is known. This means that every cell division has been tracked so we know exactly where each of the 959 cells have come from and what it does. Not all organisms have a fixed cell lineage like this, which makes C. elegans development easier to study. The fertilized egg divides into two cells and so on to generate all the cells needed to make up the worm. Some of the cells in the lineages are programmed to die. The same cells always die at the same time in development. This phenomenon of programmed cell death or cell suicide is called apoptosis and is not the same as death due to injury or damage. Apoptosis is used as a way to clear superfluous, unwanted cells and is not unique to C. elegans. C. elegans actually generates 1090 cells during its development. It ends up with 959 cells because 131 are programmed to die. We can actually see cells dying. In this photo, the arrow points to a cell about to divide. One of its daughters will undergo programmed cell death. The cell from the previous photo has divided, and the arrows point to the two daughter cells. The division was unequal, and the smaller cell on the left will die. The dying cell refracts light differently and appears as a more distinct body. Neighboring cells will engulf the dead cell and get rid of the corpse. You can see the time lapse movie of this sequence of events in the AUDIO/VIDEO section of this concept. We became interested in the genes that carry out the cell death program. We called these genes cell death abnormals (ced). One in particular, ced-3, encodes a protein required for programmed cell death to occur. The ced-3 protein is a protease that actively degrades other proteins. In ced-3 mutants, cells that normally die don’t; they survive and often assume the function of their sister cells. Another gene, ced-9, encodes a protein that prevents programmed cell death in cells that should live. In other words, the ced-9 protein promotes cell survival. Mutations in ced-9 cause cells that normally live to die. The ced-3 and ced-9 proteins interact. Whenever it is present, the ced-3 protease will act on the ced-9 protein. This prevents ced-3 from killing the cells. In fact, the more ced-9 protein that is present, the better the protection against cell death. If there is enough ced-9 protein, then even cells that are supposed to die get protected against cell death. When we searched a gene database with the ced-3 and ced-9 sequences, we found two similar human genes: caspase-9 and bcl-2. These genes encode proteins with equivalent cell death functions in humans. In fact, we can replace the C. elegans ced-9 gene with human bcl-2 gene and still protect against cell death in C. elegans. Given the role of these genes in controlling cell growth and cell death, mutations in them can contribute to unchecked or unregulated growth leading to cancer. Hi, I'm Scott Lowe. I'm interested in the regulation of cell death in cancer. p53 is known to be a tumor-suppressor gene. Mice missing both copies of the p53 gene develop multiple malignant tumors. It turns out that, like the yeast rad9 gene, p53 is a checkpoint gene that monitors the state of the DNA. p53 protein levels rise after DNA damage, and the cells stop before the S stage. When DNA damage is extreme, high levels of p53 start the cell death program, and the cell undergoes programmed cell death. Again, bcl-2 acts as the protector and promotes cell survival. So, the cell death, repair and arrest pathways can be linked to tumor growth through the p53 tumor-suppressor gene. In other words, cancer can not only be thought of as unregulated cell growth leading to proliferation, cancer can also be the result of cells that can't die when they should.
single celled organisms, leland hartwell, dna polymerase, yeast cells, dna replication, dna ligase, second gap, budding yeast, okazaki fragments, all eukaryotic cells, control mechanisms, cellular components, biological problems, nuclear membrane, streches, horvitz
- ID: 16784
- Source: DNALC.DNAFTB
Related Content
16806. Biography 38: Leland Hartwell (1939 - )
Lee Hartwell was one of the first to use yeast as a model system, and he identified many of the genes involved in the cell cycle.
16834. Animation 40: Living things share common genes.
Mike Wigler shows how all organisms share similar genes, called homologs.
16087. Animal cell
Organelles in a typical animal cell.
16877. Cell Signals
Journey inside a cell as you follow proteins and learn about cellular interactions. This 3-D animation brings to life the inner workings of a fibroblast cell as it responds to external signals. Created by Cold Spring Harbor Laboratory and Interactive Know
16705. Animation 34: Genes can be moved between species.
Stanley Cohen and Herbert Boyer transform bacteria with a recombinant plasmid, and Doug Hanahan studies induced transformation.
941. Hallmarks, Evading death
Professor Robert Weinberg discusses how cancer cells have to learn how to avoid the process of programmed cell death known as apoptosis carried out in normal cells.
16647. Higher cells incorporate an ancient chromosome.
DNAFTB Animation 30: Ivan Wallin presents his idea that mitochondria and chloroplasts were once free-living organisms.
999. Causes, Viruses: HPV
In this section learn how viruses contribute to cancer development.
548. Model Center
Model organisms share with humans many key biochemical and physiological functions that have been conserved (maintained) by evolution.
960. Causes, Smoking: p53
This series of animations shows how mutations in the p53 gene are found in 70% of lung tumors, the highest rate for any cancer.